New Method Allows Researchers to “Count” Fish Using a Glass of Water
OutdoorHub Reporters 01.16.14
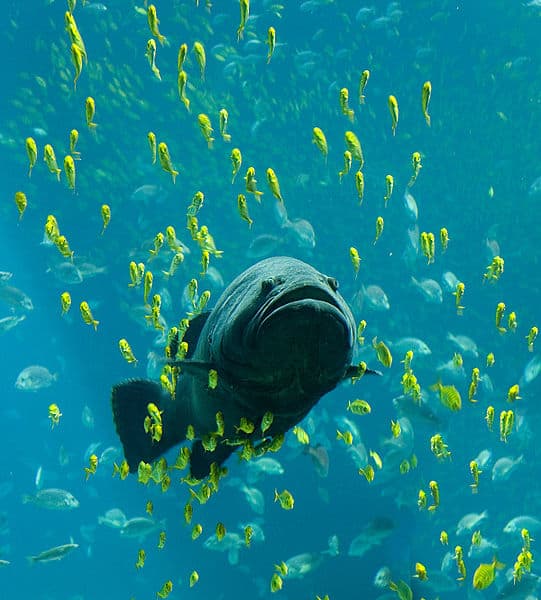

Scientists have long looked for a faster, more efficient way of counting fish in large bodies of water. This is especially true with mounting pressure from invasive species such as Asian carp and northern snakehead, making the relatively new field of DNA sampling a valuable tool in preserving America’s lakes and rivers. In a study published in the Journal PlOS ONE on Wednesday, researchers say they have reached the next step in DNA identification techniques. According to the University of Washington (UW), this method would require as little as a glass of water.
“It might be unpleasant to think about when going for a swim in the ocean, but the water is a soup of cells shed by what lives there,” said Ryan Kelly, an assistant professor at UW and the lead author of the study.
Figuring out what kinds of fish live in a particular lake is critical to proper management by wildlife officials. In the past, discerning fish species and numbers can be time-consuming and at times, amounts to guesswork. Traditional methods of counting fish often involve netting, surveys, and other sampling techniques that allow biologists to make an educated guess. Oftentimes these estimates are highly accurate, but for very small populations, traditional methods do not work. In the case of Asian carp, where even a few fish could establish a colony, biologists turn to DNA sampling.
Kelly’s method involves using molecular probes called “primers” to distinguish between different species. Researchers tested the technique on the Monterey Bay Aquarium’s Open Sea tank. Holding more than 1.2 million gallons of water, the aquarium is one of the largest in the world and contains an abundance of fish species. Kelly and his team were successful in isolating the eight bony fish species in the tank and their abundance, although the method did not work for other creatures such as turtles or sharks.
“Clearly this is an effective tool in the wild when you know what you’re looking for,” Kelly said.
Researchers involved with the study say this new method is much quicker and less expensive than traditional methods, especially with the prevalence of DNA techniques used today. A $5,300 gene sequencing test in 2001 would cost just about six cents today. Kelly’s method takes it a step further by lowering the required material needed for analysis. In testing the Open Sea tank, researchers took only about two pint glasses of water.
“More efficient, more cost-effective, and more sensitive methods could thus revolutionize ecosystem assessments and improve the way in which we collect baseline ecological data about marine ecosystems and frame monitoring efforts,” researchers wrote in the published study.